Scopus
Converting Nataliya's CSV data from Scopus into networks I noticed that the records contain besides the author name also its Scopus Author Identifier (and for journals their ISSNs).
- Jeroen Baas, Michiel Schotten, Andrew Plume, Grégoire Côté, Reza Karimi: Scopus as a curated, high-quality bibliometric data source for academic research in quantitative science studies. Quantitative Science Studies (2020) 1 (1): 377–386.
- Anne-Wil Harzing: Health warning: Might contain multiple personalities. The problem of homonyms in Thomson Reuters Essential Science Indicators. Scientometrics 105(3):2259-2270.
This provides a (at least partial) solution to the author's (and journal's) identification problem.
Data from Scopus
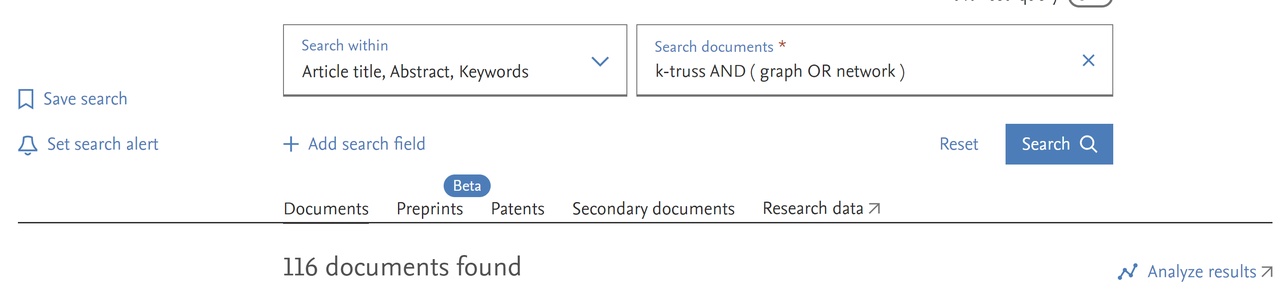
On the computer from the university/research domain (from outside using VPN), we visit the IZUM services page and click on the option Scopus. In Scopus, we enter the query determining the works of our interest. For example, the query k-truss AND ( graph OR network )
returns the following result
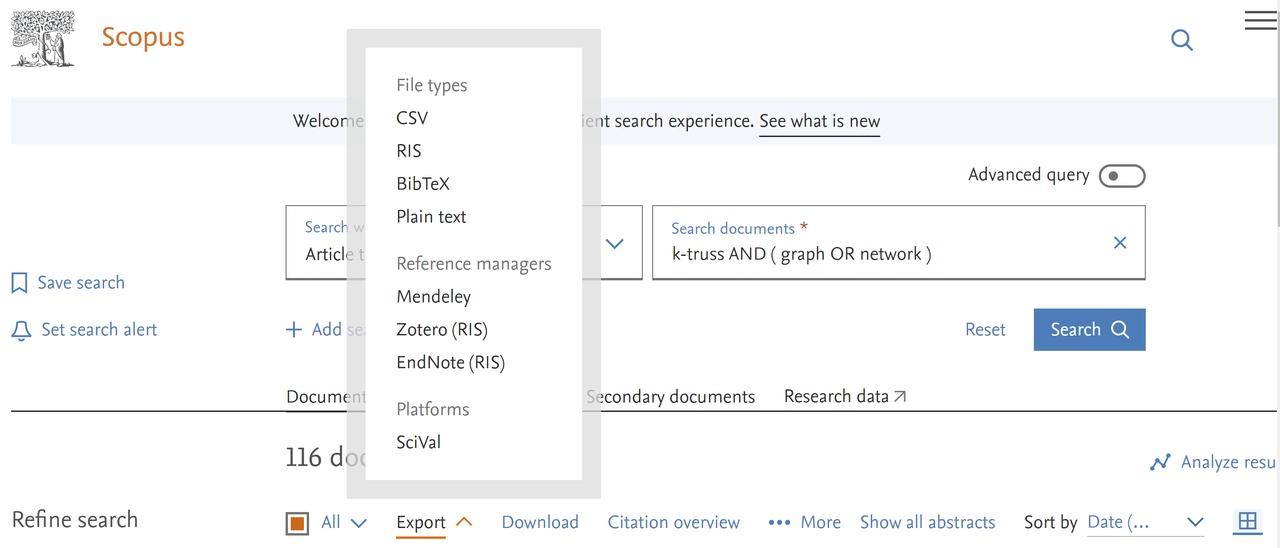
Selecting all hits (red square) and the option export we get
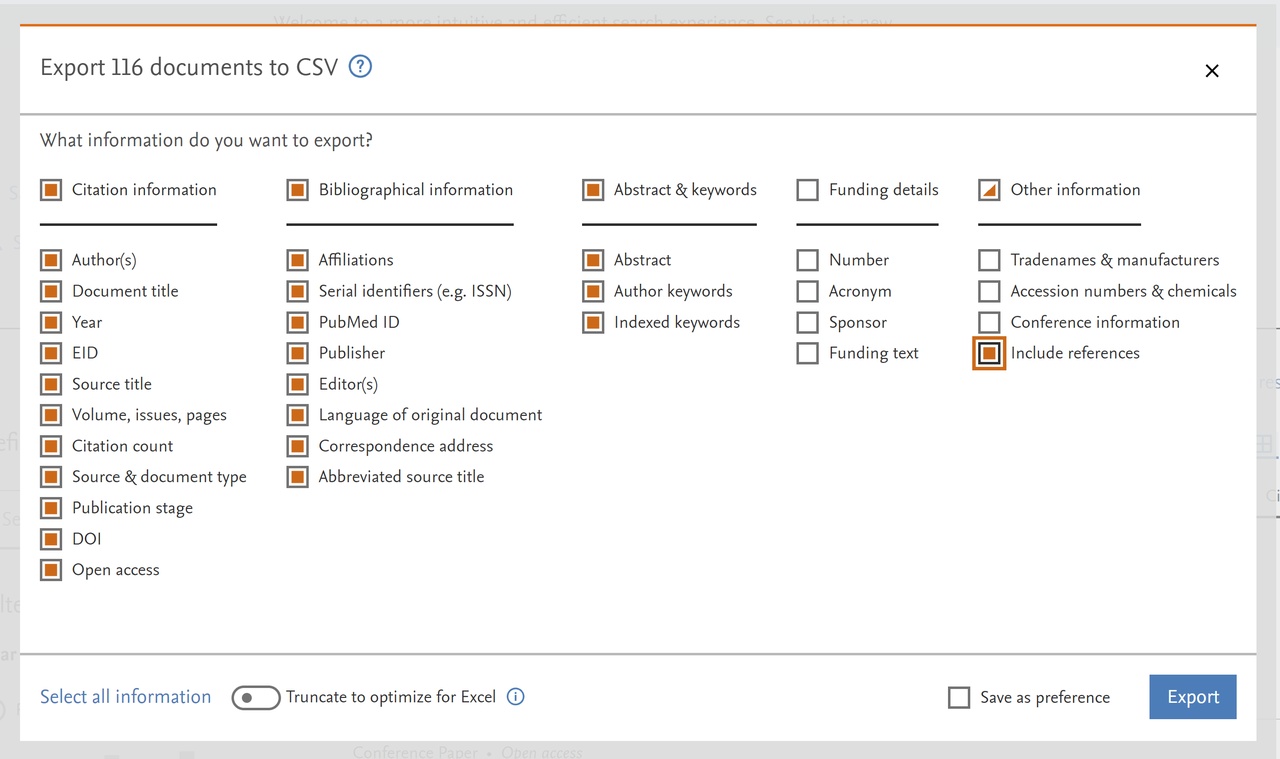
and afterward selecting the CSV option we get a window for selecting exported fields. We select all options in the first three columns, the Include references option in the last column, and switch off the option Truncate to optimize for Excel.
Clicking on the button Export we start the creation of the CSV file with our hits. In a similar way, we exported the hits also in RIS, BiBTeX, and plain text format. They are all available in the scopus-truss.zip file.
Up to 20000 hits can be saved at once.
To do: Scopus2Pajek
Because Scopus provides besides the names of units (authors, journals, works) their IDs it simplifies the identification problem for them. We would need a program Scopus2Pajek similar to WoS2Pajek (GitHub/Bavla WoS2Pajek, Slides, Python) for converting Scopus data into Pajek network files - a combination of WoS2Pajek and solutions in R.
There exists a program Scopus2WoS for the conversion of Scopus data in RIS format into WoS format. We selected the RIS format because it is very close to the WoS format. Unfortunately, it was a bad decision - in the RIS file, the IDs are not included.
In the conversion, we encountered the problem of replacing nonASCII characters with ASCII characters.